Cancer Type-Specific Mutations
Most cell lines in the public collection derive from tumor cells. In many cases the tumors exhibit type-specific mutations which contribute to tumorigenesis and are, therefore, relevant for diagnostic studies and research. These genetic markers may be identified by specific assays, informing both characterization and authentication of the cell lines.
Cancer is a disease where the malignant cells have acquired multiple mutations in genes deregulating proliferation, survival and/or differentiation. The mutated genes encode general regulators which may be altered in many tumors or in specific types of tumor only.
Tumor cell lines serve as tumor specific models for research and as positive controls in diagnostic applications. They retain many characteristics of the primary tumor, including specific gene alterations. Therefore, the confirmation of tumor/cell line specific mutations, e.g. certain fusion genes, supports the authentication of cell lines which is a prerequisite for diagnostic analyses and cancer research with cell line models.
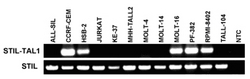
PCR is a very specific and rapid method for detecting particular gene configurations. RT-PCR analyzes the transcripts of tumor cell lines and detects the expression of, notably, fusion genes. The figure above shows an agarose gel with RT-PCR products of the gene fusion STIL-TAL1 which originates by the recurrent chromosomal aberration del(1)(p33) in acute T-cell leukemia.
Note: We use gene designations as suggested by the HUGO Gene Nomenclature Commitee (www.genenames.org). Where appropriate we also add older, perhaps better known gene names.
References:
1. van Dongen JJ, Macintyre EA, Gabert JA, Delabesse E, Rossi V, Saglio G, Gottardi E, Rambaldi A, Dotti G, Griesinger F, Parreira A, Gameiro P, Diáz MG, Malec M, Langerak AW, San Miguel JF, Biondi A: Standardized RT-PCR analysis of fusion gene transcripts from chromosome aberrations in acute leukemia for detection of minimal residual disease. Report of the BIOMED-1 Concerted Action: investigation of minimal residual disease in acute leukemia. Leukemia 13: 1901-1928 (1999).
2. van Dongen JJ, Langerak AW, Brüggemann M, Evans PA, Hummel M, Lavender FL, Delabesse E, Davi F, Schuuring E, García-Sanz R, van Krieken JH, Droese J, González D, Bastard C, White HE, Spaargaren M, González M, Parreira A, Smith JL, Morgan GJ, Kneba M, Macintyre EA: Design and standardization of PCR primers and protocols for detection of clonal immunoglobulin and T-cell receptor gene recombinations in suspect lymphoproliferations: report of the BIOMED-2 Concerted Action BMH4-CT98-3936. Leukemia 17: 2257-2317 (2003).
3. Nagel S, Venturini L, Meyer C, Kaufmann M, Scherr M, Drexler HG, MacLeod RA: Multiple mechanisms induce ectopic expression of LYL1 in subsets of T-ALL cell lines. Leukemia Res. 34: 521-528 (2010).